In this post we will compare the performance of 4 different methods to loop over a Pandas dataframe and create a new column from the data. We will compare the iterrows, itertuples and simply loop through numpy arrays. After this we will explain how to achieve an order of magnitude speed up using numba.
Problem Intro
First let's formulate a toy problem. Say we have a data frame that looks as follows
import pandas as pd
import numpy as np
import numba as nb
import time
import matplotlib.pyplot as plt
np.random.seed(0)
df = pd.DataFrame()
df['col1'] = np.random.normal(size=100000)
df['col2'] = np.random.normal(size=100000)
df['col3'] = np.random.normal(size=100000)
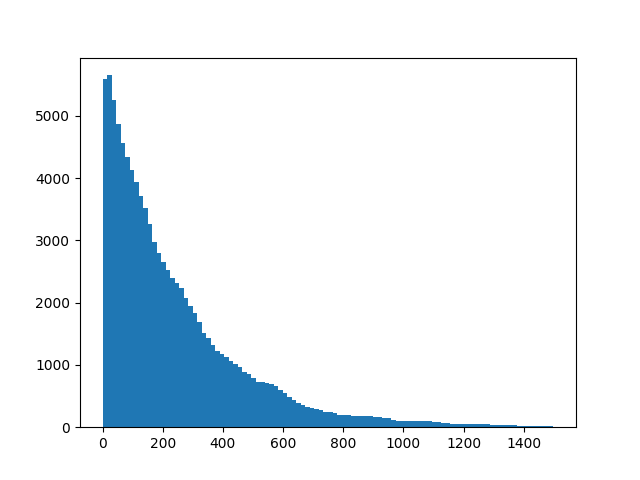
The dataset consists of 3 columns of random normal variables, each with 100k rows. We are interested in finding the distribution of waiting times between +3 sigma moves in any of the 3 columns. An event is triggered if any of the 3 rows has a value larger than 3. Then we want to store the results in a new array for later inspection. Let's do a quick sanity check which should illustrate the problem more clearly.
for i, row in df.iterrows():
if (row >3).any():
print(row)
break
'''
col1 0.906045
col2 3.264884
col3 -2.660591
Name: 111, dtype: float64
'''
Looks like we found the first value of interest after 111 iterations of the loop. We would like to do this for all rows and keep a running count of the number of periods between the events (3 sigma changes).
Note that after creating this toy examples below I realized I introduced some bias to this analysis by initializing the counter at 0. I guess if you wanted to be more rigorous in a real analysis you might want to set this to a value like -1000000000 and then replace with pd.Na after it is done, but since this example is more to demonstrate speed differentials I take view that it is clearer to set it to 0 to understand the problem at hand. There we go I just saved someone a lengthy email (yes you).
Pandas Iterrows Method
First let's try to understand the iterrows methods and why it is usually the slowest option.
for i, row in df.iterrows():
print(i, type(i))
print(row, type(row))
break
'''
0 <class 'int'>
col1 1.764052
col2 -0.483797
col3 0.039510
Name: 0, dtype: float64 <class 'pandas.core.series.Series'>
'''
As can be seen from the first row of data above, the iterrows method returns a tuple containing the row index along with a pd.Series object with the columns as indices and the values in a column. Clearly there is a lot of overhead instantiating a series object on each iteration of the loop.
def pandas_itterows(df):
res = np.empty(df.shape[0], dtype='int64')
counter = 0
for i, row in df.iterrows():
if (row > 3).any():
res[i] = counter
counter = 0
else:
counter += 1
res[i] = counter
return res
t = time.time()
res_iters = pandas_itterows(df)
print(f"pandas itterrows = {time.time()-t}")
'''
pandas itterrows = 26.616524934768677
'''
Well there is no real surprise there, we expected this to take quite a long time.
Pandas Itertuples Method
The itertuples method creates a a generator which yields a named tuple for each row of the dataframe. Let's take a look
for row in df.itertuples():
print(row, type(row))
print(row.col1, row.col2, row.col3)
break
'''
Pandas(Index=0, col1=1.764052345967664,
col2=-0.48379749195754734, col3=0.039509896265321955) <class 'pandas.core.frame.Pandas'>
1.764052345967664 -0.48379749195754734 0.039509896265321955
'''
We can access elements by name in this method. Let's calculate the waiting times and ensure the results agree with the ones we got from iterrows.
def pands_itertups(df):
res = np.empty(df.shape[0])
counter = 0
for row in df.itertuples():
i = row.Index
if row.col1 > 3 or row.col2 > 3 or row.col3 > 3:
res[i] = counter
counter = 0
else:
counter += 1
res[i] = counter
return res
t = time.time()
res_tups = pands_itertups(df)
print(f"pandas itertuples = {time.time()-t}")
print(f"agrees with iterrows {np.allclose(res_iters, res_tups)}")
'''
pandas itertuples = 0.29600954055786133
agrees with iterrows True
'''
Just by changing to the itertuples method we achieve a significant speed up, this is to be expected as there is less over-head instantiating a named tuple than creating a series object on each iteration. And thankfully we get the same results from this method as we do from itertuples.
Pure Numpy Arrays
In this method we simply convert the columns to numpy arrays and then loop over the values.
def res_numpy_arrays(df):
col1 = df.col1.to_numpy()
col2 = df.col2.to_numpy()
col3 = df.col3.to_numpy()
res = np.empty(df.shape[0])
counter = 0
for i in range(col1.shape[0]):
if col1[i] > 3 or col2[i] > 3 or col3[i] > 3:
res[i] = counter
counter = 0
else:
counter += 1
res[i] = counter
return res
t = time.time()
res_arrs = res_numpy_arrays(df)
print(f"array method = {time.time()-t}")
print(f"agrees with other methods {np.allclose(res_arrs, res_tups)}")
'''
array method = 0.15300202369689941
agrees with other methods True
'''
Again there should be no real surprise here that this method is faster than iterrows and itertuples as we avoid the overhead of instantiating objects on each iteration.
Numba Method
Here we will use eager compilation to create the waiting time method. Note that the signature (types) provided in the function below only relate to the input types i.e. 1d arrays of floating point numbers, if we leave the return type blank numba will infer it for us!
@nb.njit((nb.float64[:], nb.float64[:], nb.float64[:]))
def do_it_numba(col1, col2, col3):
res = np.empty(col1.shape[0], dtype='int64')
counter = 0
for i in range(col1.shape[0]):
if col1[i] > 3 or col2[i] > 3 or col3[i] > 3:
res[i] = counter
counter = 0
else:
counter +=1
res[i] = counter
return res
t = time.time()
res_numba = do_it_numba(df.col1.to_numpy(), df.col2.to_numpy(), df.col3.to_numpy())
print(f"numba method = {time.time()-t}")
print(f"agrees with other methods {np.allclose(res_arrs, res_numba)}")
'''
numba method = 0.0010442733764648438
agrees with other methods True
'''
It looks like numba took just 1 millisecond to complete the task, without doing proper speed tests, it is pretty clear that numba is the clear winner, and results in a significant speed up.
Let's try a speed-test with a larger dataframe, because if you are interested in speeding something up, it is unlikely the difference we have seen so far will even matter. Below we try a larger example on 10 million rows.
df = pd.DataFrame()
df['col1'] = np.random.normal(size=10000000)
df['col2'] = np.random.normal(size=10000000)
df['col3'] = np.random.normal(size=10000000)
%timeit res = do_it_numba(df.col1.to_numpy(), df.col2.to_numpy(), df.col3.to_numpy())
'''
93.6 ms ± 7.06 ms per loop (mean ± std. dev. of 7 runs, 10 loops each)
'''
%timeit res_arrs = res_numpy_arrays(df)
'''
16.9 s ± 1.83 s per loop (mean ± std. dev. of 7 runs, 1 loop each)
'''
It looks like we have achieved a speedup somewhere in the region of 170x when compared to the fastest non-numba method we have tested.
And there we have it, we have our waiting times, without having to wait too long, thanks to numba.
plt.hist(res_numba, bins=100)
Summary
- Pandas iterrows method is very slow indeed. And should be avoided where possible. Although, it is also true that there is a lot of convenience associated with using this method especially if we want to use the built-in methods.
- Pandas itertuples is a nice alternative if we want to be able to access variables by name. It is significantly faster due to the generator express using named tuples rather than pd.Series objects.
- Numba is so incredibly fast!! Where possible we should consider switching anything that involves looping over pandas dataframes or columns to using Numba.